Choosing K and the denoising parameters
Source:vignettes/choosing-k-denoising.Rmd
choosing-k-denoising.RmdThis vignette focuses on two practical knobs in the MetaHunt
pipeline: the latent rank K and the d-fSPA denoising
parameters (N, Delta). For the broader setup — the four
assumptions, the three-step pipeline, and the running notation — see
vignette("metahunt-intro", package = "MetaHunt").
Why this matters
Choosing K is the single most consequential decision in
a MetaHunt fit. Picking K too small underfits: real
cross-study heterogeneity gets squashed into a low-rank approximation
that cannot represent the data, and downstream predictions are biased.
Picking K too large inflates variance and risks recovering
spurious “bases” that fit noise. The denoising step in d-fSPA controls
finite-sample variance in a complementary way: averaging each study with
its near neighbours before basis hunting smooths over per-study
estimation error, at the cost of a small smoothing bias.
A small standalone simulation
m <- 30; G <- 20; K_true <- 3
x <- seq(0, 1, length.out = G)
basis <- rbind(sin(pi * x), cos(pi * x), x)
W <- data.frame(w1 = rnorm(m), w2 = rnorm(m))
beta <- cbind(c(1, -0.8), c(-0.5, 1.2), c(0, 0))
pi_true <- exp(as.matrix(W) %*% beta); pi_true <- pi_true / rowSums(pi_true)
F_hat <- pi_true %*% basis + matrix(rnorm(m * G, sd = 0.05), m, G)Unsupervised diagnostic: reconstruction error vs K
The elbow plot tracks how well the recovered bases
reconstruct the observed F_hat as a function of
K. It is unsupervised — it does not use W —
and is fast.
elbow <- reconstruction_error_curve(F_hat, K_range = 2:6,
dfspa_args = list(denoise = FALSE))
plot(elbow$K, elbow$error, type = "b",
xlab = "K", ylab = "reconstruction error",
main = "Reconstruction error vs K",
ylim = c(0, max(elbow$error, na.rm = TRUE) * 1.05))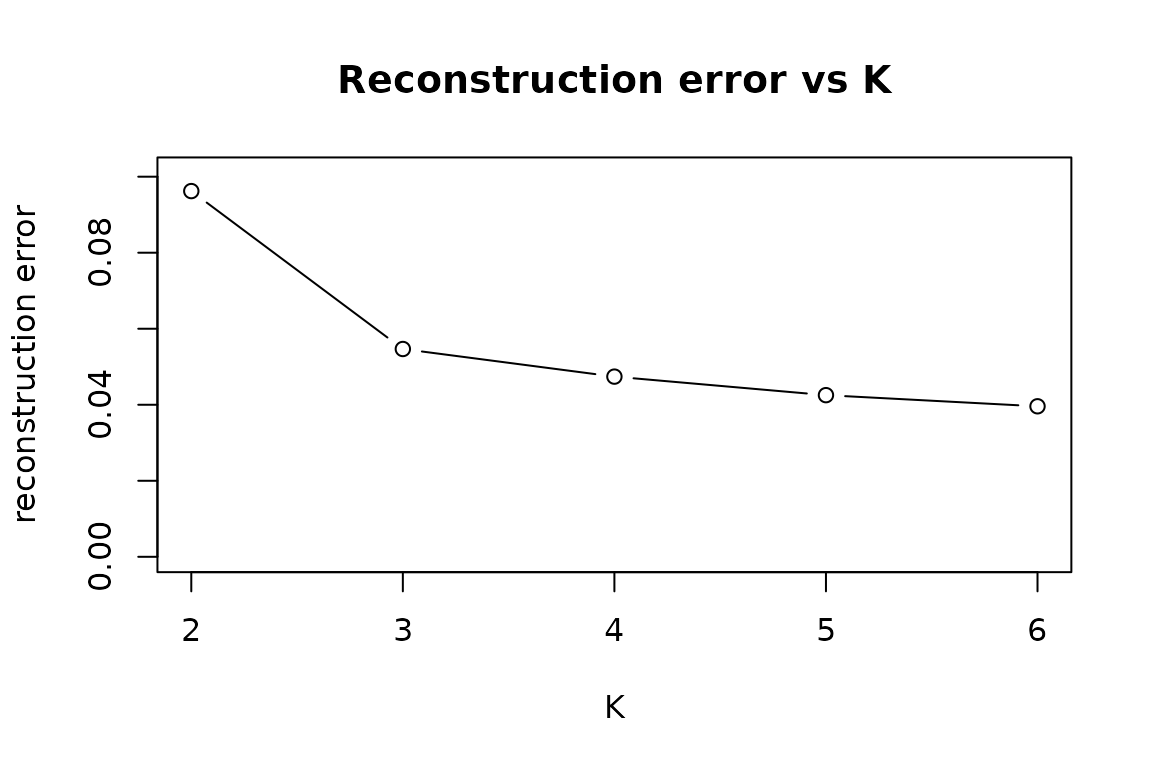
Supervised diagnostic: cross-validated prediction error vs K
The CV prediction-error curve uses the metadata
W to predict held-out studies’ functions and reports the
average prediction error. This is supervised and tends to identify a
tighter elbow when the metadata is informative.
cv <- cv_error_curve(F_hat, W, K_range = 2:6, n_folds = 4,
dfspa_args = list(denoise = FALSE), seed = 1)
plot(cv$K, cv$cv_error, type = "b",
xlab = "K", ylab = "CV prediction error",
main = "CV prediction error vs K",
ylim = c(0, max(cv$cv_error, na.rm = TRUE) * 1.05))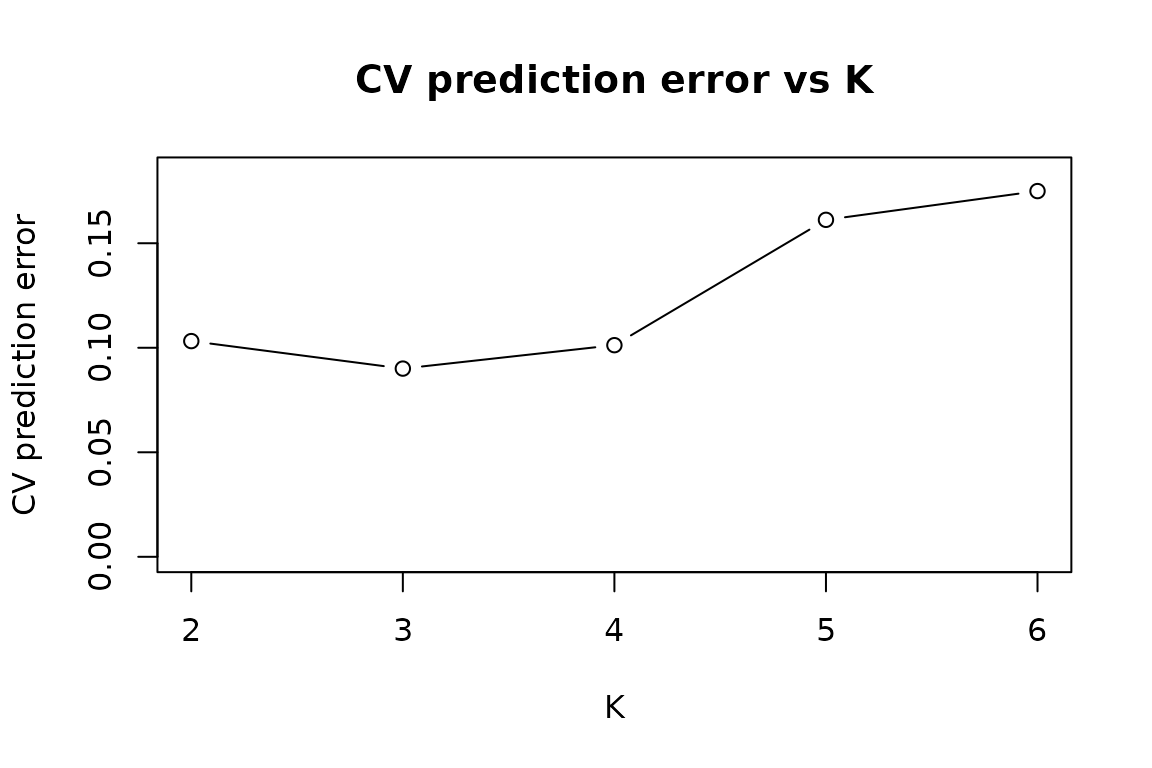
Both curves should dip near K = 3, the true rank in this
simulation.
The d-fSPA denoising knobs (N, Delta)
dfspa() averages each study with its near neighbours
before running the projection algorithm. Two parameters control this:
N (the neighbourhood size, in number of studies) and
Delta (a distance threshold). Larger N and
Delta smooth more aggressively.
Bypassing denoising
In clean simulations or with small m, the simplest
choice is to bypass denoising entirely. This avoids the small-sample
failure mode where aggressive denoising prunes too many studies.
Setting (N, Delta) by hand
If you have a sense of scale for the within-study estimation error,
pass N and Delta directly. These two calls
illustrate a hand-tuned and a near-default configuration on the same
data.
Tuning (N, Delta) by CV
select_denoising_params() cross-validates over a grid of
(N, Delta) combinations at fixed K. With small
m, the search will frequently warn that some combinations
prune everything (“Only 0 studies survive denoising but K = 3…”). These
warnings are expected: aggressive (N, Delta) on small
training folds is too strong. The function records those folds as
failures and returns the best surviving combination.
tune <- select_denoising_params(F_hat, W, K = K_true, n_folds = 4, seed = 1)
tune$best
#> $N
#> [1] 0.6802395
#>
#> $Delta
#> [1] 0.04555978
#>
#> $cv_error
#> [1] 0.0900349Practical recipe
- Start with the elbow plot to get a rough range for
K. Refine with the CV curve ifWis informative. - For very small
m(saym < 30), bypass denoising (denoise = FALSE) and pickKfrom the CV curve. - For larger
m, leave the d-fSPA defaults on or tune(N, Delta)withselect_denoising_params(). - Treat warnings from
select_denoising_params()as informative, not fatal. The reportedbestis the best surviving combination. - Sanity-check the recovered bases visually with
plot(fit). Bases that look like noise are a sign ofKset too high.
See also
-
vignette("metahunt-intro", package = "MetaHunt")— the full pipeline and key assumptions. -
?metahunt— the wrapper around the three pipeline steps. -
?dfspa— d-fSPA basis hunting and its denoising arguments. -
?reconstruction_error_curve— the unsupervised elbow diagnostic. -
?cv_error_curve— the supervised CV diagnostic. -
?select_denoising_params— cross-validating(N, Delta)at fixedK.